5. Agora Data Model¶
The Agora Data Model is implemented along two different lines. On the one side, arbitrary files embedded in an arbitrary directory structure are represented in the project folder tree. Based on the file name extension, Agora may be able to display some files, such as jpeg image files with the image viewer or text files in a text viewer. On the other side, Agora also implements a Data Model based on a simplified Dicom Model of the Real World. In this model, data acquired on a patient in a single session is organized in a study. A study contains an arbitrary number of series, where one series typically contains the data from one acquisition. This data consists of an arbitrary number of datasets, where the datasets represent the same data indifferent form (such as raw data and reconstructed image data) or data associated with each other (such as image data and respective logging data). A dataset finally is composed of one or several data files. Studies, series and datasets are Agora data model objects and referred as Agora objects in this manual.
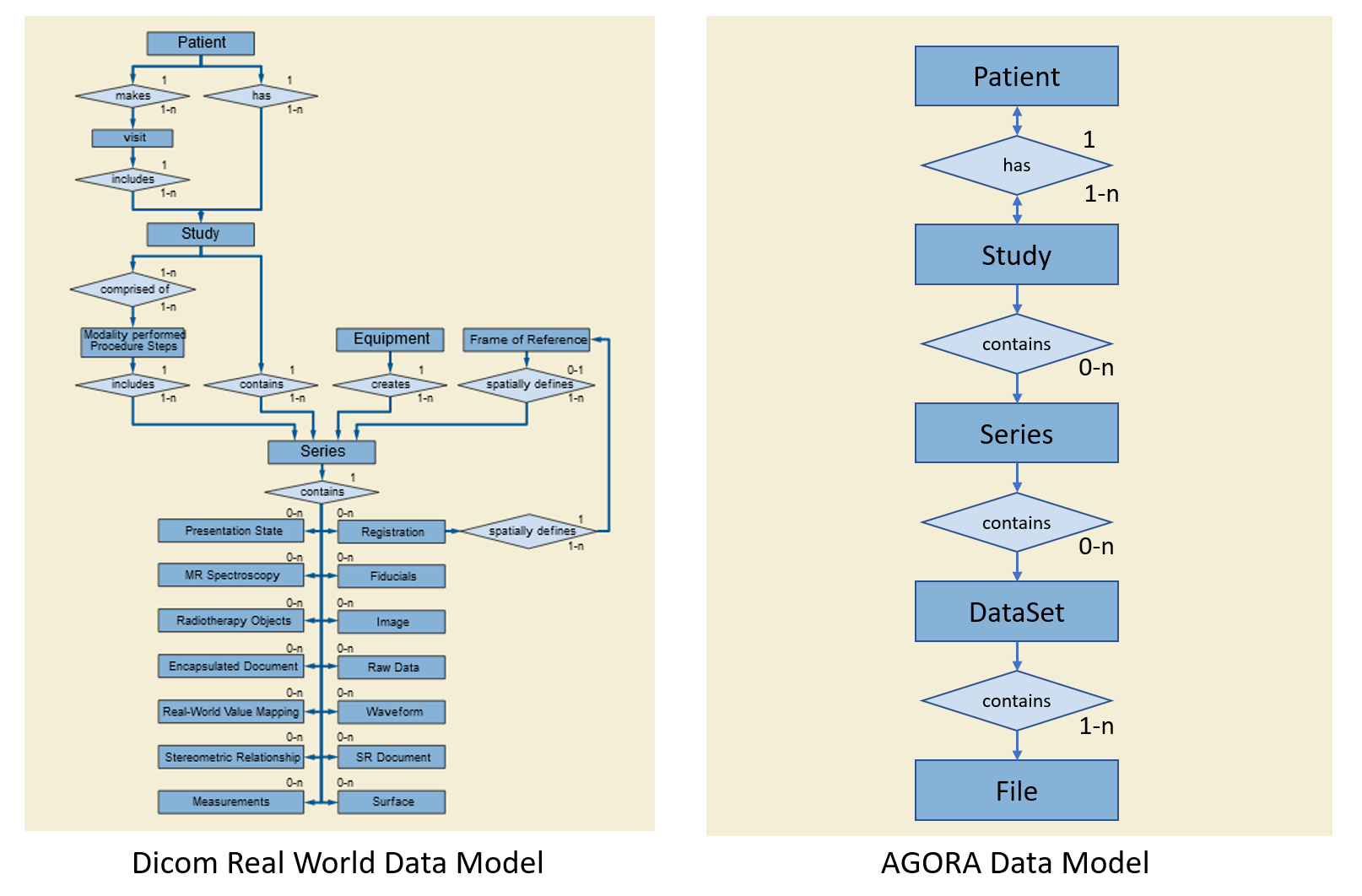
Data that fits the Agora data model is imported as one or multiple studies in Agora. The data appears in the project study list only, and in particular not in the project folder tree. It is however possible to insert a reference to a study (or series or dataset) into the folder tree. The same Agora object can be referenced multiple times in the project folder tree. The actual data is not copied, all references point to the same files.
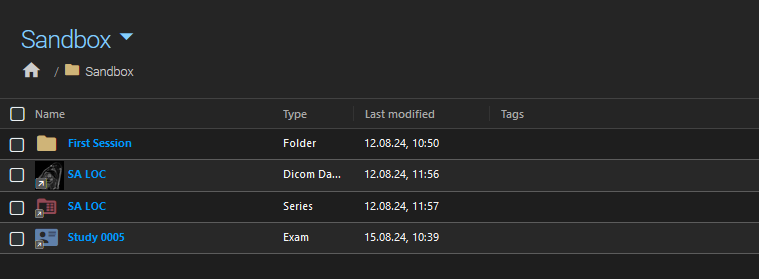
Example of a folder containing references to study, series, and dataset.¶
5.1. Parameter Sets¶
Also part of the Agora Data Model are the parameter sets. These are list of parameters associated to an Agora object and stored in the Agora database. Typically meta-data such as the Dicom tags in Dicom files are extracted from imported files and stored into parameter sets.

A parameter set containing Dicom tag values and associated to an AGORA dataset.¶
In addition to these automatically extracted parameters, Agora also supports user-defined parameters — custom key/value metadata that you can attach manually to studies, series, patients, and folders via the Parameters tab in the detail panel. User-defined parameters are stored in the same parameter set structure and are searchable using the same AQL query syntax. See User-Defined Parameters in the Managing Data chapter for details.
5.2. Data Upload vs. Data Import¶
When uploading data files to Agora, files that do not match the Agora data model and for which no parameters have been specified, will be stored in the project folder tree (in the root folder if no destination folder is specified). Files that do match the Agora data model will be imported as a new or in an existing study. In addition, metadata available in the files (depending on the file type) will be extracted and stored as parameter sets in the Agora database. When uploading file collections to Agora, files matching the Agora data model will be extracted from the collection and imported as Agora objects (studies). The remaining files of the collection will be uploaded into the folder tree. Agora will attempt to insert object references at the folder tree locations the object files were extracted from.
5.3. Segmented Data Import¶
Agora data model objects (studies, series, datasets) are labelled with unique identifiers. Child objects hold a reference to the identifier of their parent object. This allows for the segmented import of data. A partial import of a study can be completed at a later time. Agora checks for already existing parent identifiers and if given will import into the existing parent. Importing objects already existing in a project will be ignored.
Example: A scanning session produces ten series, but only five are transferred to Agora today. Agora creates the study and imports those five series. The following day the remaining five series are transferred. Because they carry the same study UID as the first batch, Agora recognises the existing study and adds the missing series to it — no duplicate study is created and no data from the first import is touched.
5.4. Data Storage and Project Isolation¶
All Agora objects — patients, studies, series, and datasets — are project-scoped. When data is copied from one project to another, new independent instances of each object are created in the destination project. The same study appearing in two projects has no shared state: renaming, deleting, or modifying it in one project has no effect on the other.
This project isolation also applies to data that is imported into a project where it technically already exists in another project. Agora does not detect cross-project duplicates at the object level; each project receives its own independent copy.
5.4.1. File-Level Sharing (Copy-on-Write)¶
The one exception to strict project isolation is at the file level. The actual data files on disk (represented by the DataFile model) are shared between datasets using a many-to-many relationship. When a dataset is copied, the new dataset object references the same physical files as the original — no disk space is consumed for the file data itself.
A physical copy of a file is only made when the file needs to be modified. Currently the only operation that triggers such a copy is anonymization: when a dataset is anonymized, Agora creates a new copy of each affected file with the personal data removed, and the dataset is updated to reference the new files. The original files remain on disk for as long as any other dataset still references them.
This copy-on-write approach means that storing many copies of the same data across different projects costs little extra disk space, while each project’s data remains fully independent at the object level.
5.5. Duplicate Import Behaviour¶
When data is imported into a project where part or all of that data already exists, Agora uses the following rules to decide what to create and what to skip.
5.5.1. Study and Series Merging¶
Studies and series carry unique identifiers (UIDs derived from the scanner or the import manifest). When an import arrives:
If a study with the same UID already exists in the same project, the incoming data is merged into the existing study. New series and datasets are added; existing ones are updated or skipped.
If the same UID exists in a different project, a new independent study is created in the target project (a warning is recorded in the import log).
If no matching UID is found, a new study is created.
The same logic applies to series within a study: a series with a matching UID is reused; otherwise a new series is created. For Philips data, the acquisition number is used as a fallback identifier when the UID is unavailable.
5.5.2. Dataset and File Deduplication¶
At the dataset level, Agora uses content hashes to detect duplicates. Before checking for duplicates, Agora computes a SHA1 hash and records the size of each incoming file. It then compares the set of (type, hash, size, filename) tuples for the incoming dataset against all existing datasets in the target series or study.
If the incoming dataset’s file set exactly matches an existing dataset (same hashes, sizes, and filenames in any order), the import is skipped — the dataset is considered already present. If even one file differs, a new dataset is created.
This means:
Importing the exact same files twice into the same series is a no-op on the second import; the existing dataset is kept and nothing is overwritten.
Importing a modified version of a file (different content, same filename) creates a new dataset alongside the existing one.
Importing the same files into a different project always creates a new dataset object, but the physical files on disk are shared (see File-Level Sharing (Copy-on-Write) above).
5.5.3. Special Cases¶
Raw datasets (Philips Raw, Siemens Raw, Bruker Raw, ISMRMRD): a series may contain at most one raw dataset. If a raw dataset is already present, any subsequent raw dataset in the same import is skipped.
Bruker subject datasets: only one subject-level dataset is allowed per study. Additional subject datasets in the same import are ignored.